|
Size: 2112
Comment:
|
← Revision 8 as of 2019-02-19 18:20:02 ⇥
Size: 2970
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 9: | Line 9: |
| 1. Download the desired registration markers and bone STLs (place in folder named mri-oks00x, where x is the specimen ID). * 3 registration marker stls for tibia * 3 registration marker stls for femur * stl for tibia bone * stl for femur bone 1. Create registration xml ([[https://simtk.org/svn/openknee/utl/Registration/Registration_final/oks001_registration_01.xml| Example Registration XML]]) |
1. Download the desired registration markers and bone STLs (place in folder named geometry-oks00x, where x is the specimen ID). * For tibiofemoral joint mechanics: * 3 registration marker stls for tibia * 3 registration marker stls for femur * stl for tibia bone (only needed for visualization) * stl for femur bone (only needed for visualization) * For patellofemoral joint mechanics (in addition to TF joint mechanics): * 3 registration marker stls for patella * stl for patella (only needed for visualization) 1. Add digitizing order of femur and tibia in a [[https://simtk.org/svn/openknee/utl/Registration/Registration_final/ReadMe_Notes.txt|ReadMe_Notes.txt]] file within the main Configuration folder of the tibiofemoral test directory. 1. Create registration xml ([[https://simtk.org/svn/openknee/utl/Registration/Registration_final/oks001_registration_01.xml| Example Registration XML]]). Place in registration-00x_xx folder within the registration-00x directory. |
| Line 18: | Line 23: |
| * Inputs: * Registration xml * Boolean: 0-does not show visualization figure, 1-shows visualization figure * Outputs: * Transformation matrices saved within .npz file for each trial that has processed data. |
Target
- Create visualization of experimental tests in modeling (image) coordinate system.
- Prepare experimental data for model inputs.
Registration
- Download joint mechanics .zip file from Downloads page.
- Download the desired registration markers and bone STLs (place in folder named geometry-oks00x, where x is the specimen ID).
- For tibiofemoral joint mechanics:
- 3 registration marker stls for tibia
- 3 registration marker stls for femur
- stl for tibia bone (only needed for visualization)
- stl for femur bone (only needed for visualization)
- For patellofemoral joint mechanics (in addition to TF joint mechanics):
- 3 registration marker stls for patella
- stl for patella (only needed for visualization)
- For tibiofemoral joint mechanics:
Add digitizing order of femur and tibia in a ReadMe_Notes.txt file within the main Configuration folder of the tibiofemoral test directory.
Create registration xml (Example Registration XML). Place in registration-00x_xx folder within the registration-00x directory.
Run script from command line Registration Script
Change directory to desired OpenKnee specimen (e.g. oks001)
Run convert_to_image_CS.py <oks00x_registration_0x.xml> <1>
- Inputs:
- Registration xml
- Boolean: 0-does not show visualization figure, 1-shows visualization figure
- Outputs:
- Transformation matrices saved within .npz file for each trial that has processed data.
- Inputs:
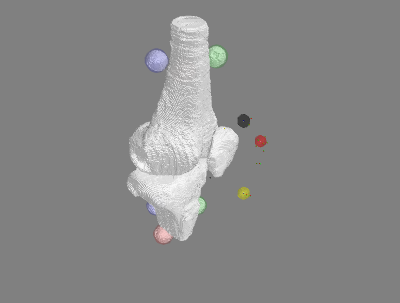
Example using oks001 - 004_passive flexion
White stls - segmentation from MRI. Colored registration marker stls - spheres from calculated center of digitized points.
Re-sample Data for Model Input
Create text file with desired tdms file(s) you would like to re-sample. Place this text file in either the KinematicsKinetics directory of the PatellofemoralJoint or TibiofemoralJoint mechanics directories depending on which data you are interested in re-sampling. Example text file for PatellofemoralJoint
Place in the appropriate directory KinematicsKinetics of either the TibioFemoral or PatelloFemoral test directories.
Run script from command line Re-sampling Script
Change directory to desired OpenKnee specimen (e.g. oks001)
Run resampling_plotting.py <oks00x_registration_0x.xml> <oks0x_xF.txt>
See Sample Passive Flexion Video for video example.