The functionality of OpenMM will (eventually) include everything that one would need to run modern molecular simulation.
-
Gideon Simpson
- Posts: 28
- Joined: Tue Sep 26, 2017 11:15 pm
Post
by Gideon Simpson » Wed May 15, 2019 12:35 pm
I am not sure if this is a genuine issue or not, but I ran some simple LJ cluster experiments with what I thought were periodic boundary conditions. The output, when viewed in Ovito, looks a bit more like shrink wrapped boundary conditions have been applied (i.e., the bounding box changes in size throughout the visualization). Perhaps this is just an artifact of how the PDB output data is pulled into Ovito.
I constructed my BCs with commands like:
Code: Select all
# other code
box_vectors = np.diag([box_edge/unit.angstrom for i in range(3)]) * unit.angstroms
system.setDefaultPeriodicBoxVectors(*box_vectors)
# other code
force = openmm.NonbondedForce()
force.setNonbondedMethod(openmm.NonbondedForce.CutoffPeriodic)
# other code
simulation.reporters.append(app.PDBReporter('output.pdb', 100,enforcePeriodicBox=True))
I'm not sure if I missed something.
-
Peter Eastman
- Posts: 2541
- Joined: Thu Aug 09, 2007 1:25 pm
Post
by Peter Eastman » Wed May 15, 2019 2:34 pm
I'm not sure what you mean by shrink wrapped boundary conditions. Does your simulation include a barostat? If so, the box will change size. Otherwise, it should stay fixed. Since you specified enforcePeriodicBox=True, the output should have all molecules translated into a single box. That is applied to the center of each molecule, so a molecule may still stick out beyond the edge of the box.
-
Gideon Simpson
- Posts: 28
- Joined: Tue Sep 26, 2017 11:15 pm
Post
by Gideon Simpson » Wed May 15, 2019 2:55 pm
I'm attaching a screen capture from Ovito. Again, this may be purely an artifact of how Ovito is interpreting the PDB output, in which case it's not really a problem. But notice that the dimensions of the cell, which started off cubic, are no longer uniform and that the locations of the particles appear to be determining the limits of the computational cell.
-
Attachments
-
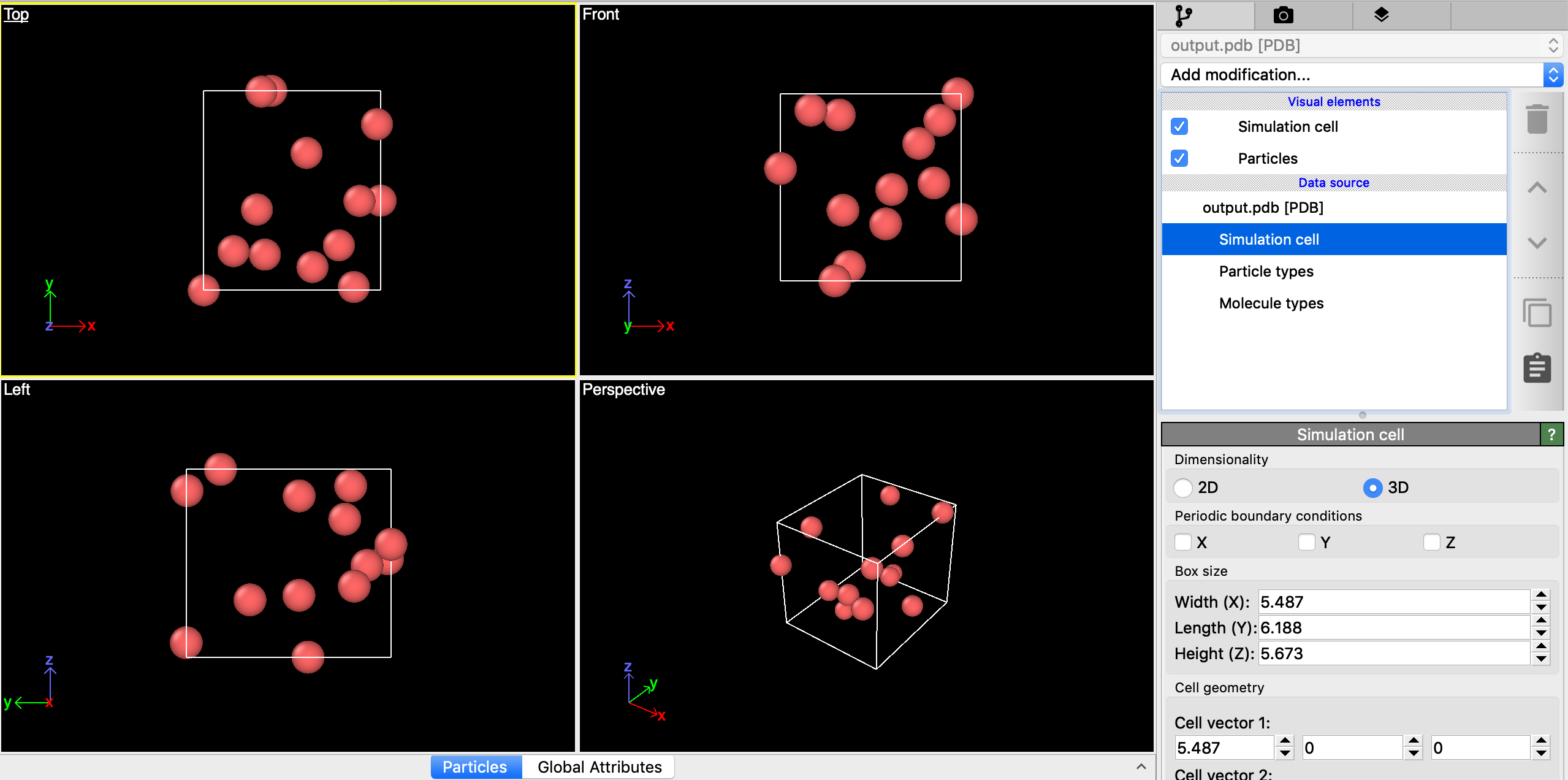
- Screen Shot 2019-05-15 at 5.52.13 PM.png (415.93 KiB) Viewed 369 times
-
Peter Eastman
- Posts: 2541
- Joined: Thu Aug 09, 2007 1:25 pm
Post
by Peter Eastman » Wed May 15, 2019 4:54 pm
That looks to me like the box it's drawing is the minimum box containing all the particles. Which is not the same as the periodic box used in the simulation.
-
Gideon Simpson
- Posts: 28
- Joined: Tue Sep 26, 2017 11:15 pm
Post
by Gideon Simpson » Wed May 15, 2019 7:13 pm
So does that mean that I'm writing PDB data incorrectly, or should I ask the Ovito people about how to post process it in the way I had anticipated?
-
Peter Eastman
- Posts: 2541
- Joined: Thu Aug 09, 2007 1:25 pm
Post
by Peter Eastman » Thu May 16, 2019 8:22 am
Neither one. It just means you should ignore the box it's drawing around the atoms. It doesn't mean what you think it does.
-
william peterson
- Posts: 1
- Joined: Thu Nov 21, 2019 4:17 am
Post
by william peterson » Thu Nov 21, 2019 4:22 am
custom writing
Neither one. It just means you should ignore the box it's drawing around the atoms. It doesn't mean what you think it does.
What's the reason it's drawing the box around the atoms? Sorry, I was confused too and didn't get this yet.
-
Peter Eastman
- Posts: 2541
- Joined: Thu Aug 09, 2007 1:25 pm
Post
by Peter Eastman » Thu Nov 21, 2019 10:18 am
Because it thinks it will be useful to you? There may be an option to turn it off. I've never used that program.